Hi Max,
thank you for your reply.
I used your link for the latest LULC example notebook (https://github.com/sentinel-hub/eo-learn/blob/master/examples/land-cover-map/SI_LULC_pipeline.ipynb) and tried it again. The Error is the same.
KeyError: “During execution of task AddValidDataMaskTask: ‘CLM’”
In the following text is the Jupyter-Notbook and the error-file:
**Jupyter-Notebook:**
# Firstly, some necessary imports
# Jupyter notebook related
%reload_ext autoreload
%autoreload 2
%matplotlib inline
# Built-in modules
import pickle
import sys
import os as os
import datetime
import itertools
from aenum import MultiValueEnum
# Basics of Python data handling and visualization
import numpy as np
np.random.seed(42)
import geopandas as gpd
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
from matplotlib.colors import ListedColormap, BoundaryNorm
from mpl_toolkits.axes_grid1 import make_axes_locatable
from shapely.geometry import Polygon
from tqdm.auto import tqdm
# Machine learning
import lightgbm as lgb
#from sklearn.externals import joblib
#from sklearn import metrics
#from sklearn import preprocessing
# Imports from eo-learn and sentinelhub-py
from eolearn.core import EOTask, EOPatch, LinearWorkflow, FeatureType, OverwritePermission, \
LoadTask, SaveTask, EOExecutor, ExtractBandsTask, MergeFeatureTask
from eolearn.io import SentinelHubInputTask, ExportToTiff
from eolearn.mask import AddMultiCloudMaskTask, AddValidDataMaskTask
from eolearn.geometry import VectorToRaster, PointSamplingTask, ErosionTask
from eolearn.features import LinearInterpolation, SimpleFilterTask, NormalizedDifferenceIndexTask
from sentinelhub import UtmZoneSplitter, BBox, CRS, DataSource,SentinelHubRequest
# Folder where data for running the notebook is stored
#DATA_FOLDER = os.path.join('..', '..', 'example_data')
DATA_FOLDER = os.path.join('D:/eoTest/example_data')
print('DATA_FOLDER: ',DATA_FOLDER)
# Load geojson file
#country = gpd.read_file(os.path.join(DATA_FOLDER, 'svn.geojson'))#svn_utm_33N
#country = gpd.read_file(os.path.join(DATA_FOLDER, 'svn_3857.geojson'))#svn_utm_33N
country = gpd.read_file(os.path.join(DATA_FOLDER, 'svn_utm_33N.geojson'))#svn_utm_33N
country = country.buffer(500)
# Get the country's shape in polygon format
country_shape = country.geometry.values[-1]
# Plot country
country.plot()
plt.axis('off');
# Print size
print('Dimension of the area is {0:.0f} x {1:.0f} m2'.format(country_shape.bounds[2] - country_shape.bounds[0],
country_shape.bounds[3] - country_shape.bounds[1]))
DATA_FOLDER: D:/eoTest/example_data
Dimension of the area is 243184 x 161584 m2
# Create the splitter to obtain a list of bboxes
bbox_splitter = UtmZoneSplitter([country_shape], country.crs, 5000)
bbox_list = np.array(bbox_splitter.get_bbox_list())
info_list = np.array(bbox_splitter.get_info_list())
# Prepare info of selected EOPatches
geometry = [Polygon(bbox.get_polygon()) for bbox in bbox_list]
idxs = [info['index'] for info in info_list]
idxs_x = [info['index_x'] for info in info_list]
idxs_y = [info['index_y'] for info in info_list]
gdf = gpd.GeoDataFrame({'index': idxs, 'index_x': idxs_x, 'index_y': idxs_y},
crs=country.crs,
geometry=geometry)
# select a 5x5 area (id of center patch)
ID = 616
# Obtain surrounding 5x5 patches
patchIDs = [616]
'''
for idx, [bbox, info] in enumerate(zip(bbox_list, info_list)):
if (abs(info['index_x'] - info_list[ID]['index_x']) <= 2 and
abs(info['index_y'] - info_list[ID]['index_y']) <= 2):
patchIDs.append(idx)
# Check if final size is 5x5
if len(patchIDs) != 5*5:
print('Warning! Use a different central patch ID, this one is on the border.')
# Change the order of the patches (used for plotting later)
patchIDs = np.transpose(np.fliplr(np.array(patchIDs).reshape(5, 5))).ravel()
'''
# save to shapefile
shapefile_name = (os.path.join(DATA_FOLDER, 'grid_slovenia_500x500.gpkg'))
# save to shapefile
#shapefile_name = './grid_slovenia_500x500.gpkg'
gdf.to_file(shapefile_name, driver='GPKG')
# figure
fig, ax = plt.subplots(figsize=(30, 30))
gdf.plot(ax=ax,facecolor='w',edgecolor='r',alpha=0.5)
country.plot(ax=ax, facecolor='w',edgecolor='b',alpha=0.5)
ax.set_title('Selected 5x5 tiles from Slovenia', fontsize=25);
for bbox, info in zip(bbox_list, info_list):
geo = bbox.geometry
ax.text(geo.centroid.x, geo.centroid.y, info['index'], ha='center', va='center')
gdf[gdf.index.isin(patchIDs)].plot(ax=ax,facecolor='g',edgecolor='r',alpha=0.5)
plt.axis('off');
class SentinelHubValidData:
"""
Combine Sen2Cor's classification map with `IS_DATA` to define a `VALID_DATA_SH` mask
The SentinelHub's cloud mask is asumed to be found in eopatch.mask['CLM']
"""
def __call__(self, eopatch):
return np.logical_and(eopatch.mask['IS_DATA'].astype(np.bool),
np.logical_not(eopatch.mask['CLM'].astype(np.bool)))
class CountValid(EOTask):
"""
The task counts number of valid observations in time-series and stores the results in the timeless mask.
"""
def __init__(self, count_what, feature_name):
self.what = count_what
self.name = feature_name
def execute(self, eopatch):
eopatch.add_feature(FeatureType.MASK_TIMELESS, self.name, np.count_nonzero(eopatch.mask[self.what],axis=0))
return eopatch
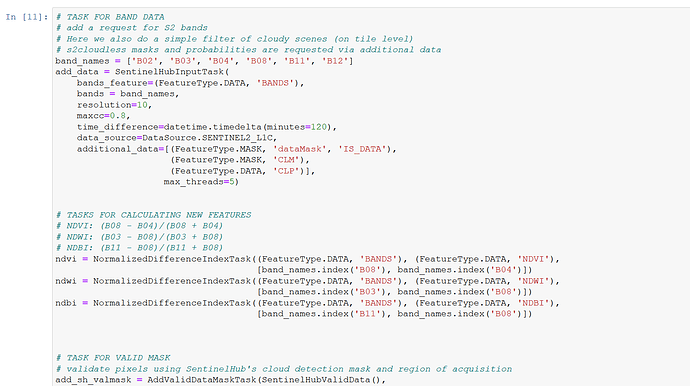
# TASK FOR BAND DATA
# add a request for S2 bands
# Here we also do a simple filter of cloudy scenes (on tile level)
# s2cloudless masks and probabilities are requested via additional data
band_names = ['B02', 'B03', 'B04', 'B08', 'B11', 'B12']
add_data = SentinelHubInputTask(
bands_feature=(FeatureType.DATA, 'BANDS'),
bands = band_names,
resolution=10,
maxcc=0.8,
time_difference=datetime.timedelta(minutes=120),
data_source=DataSource.SENTINEL2_L1C,
additional_data=[(FeatureType.MASK, 'dataMask', 'IS_DATA'),
(FeatureType.MASK, 'CLM'),
(FeatureType.DATA, 'CLP')])
# TASKS FOR CALCULATING NEW FEATURES
# NDVI: (B08 - B04)/(B08 + B04)
# NDWI: (B03 - B08)/(B03 + B08)
# NDBI: (B11 - B08)/(B11 + B08)
ndvi = NormalizedDifferenceIndexTask((FeatureType.DATA, 'BANDS'), (FeatureType.DATA, 'NDVI'),
[band_names.index('B08'), band_names.index('B04')])
ndwi = NormalizedDifferenceIndexTask((FeatureType.DATA, 'BANDS'), (FeatureType.DATA, 'NDWI'),
[band_names.index('B03'), band_names.index('B08')])
ndbi = NormalizedDifferenceIndexTask((FeatureType.DATA, 'BANDS'), (FeatureType.DATA, 'NDBI'),
[band_names.index('B11'), band_names.index('B08')])
# TASK FOR VALID MASK
# validate pixels using SentinelHub's cloud detection mask and region of acquisition
add_sh_valmask = AddValidDataMaskTask(SentinelHubValidData(),
'IS_VALID' # name of output mask
)
# TASK FOR COUNTING VALID PIXELS
# count number of valid observations per pixel using valid data mask
count_val_sh = CountValid('IS_VALID', # name of existing mask
'VALID_COUNT' # name of output scalar
)
#path_out = DATA_FOLDER
path_out = os.path.join('D:/eoTest/example_data/test')
# TASK FOR SAVING TO OUTPUT (if needed)
path_out = './eopatches/'
if not os.path.isdir(path_out):
os.makedirs(path_out)
save = SaveTask(path_out, overwrite_permission=OverwritePermission.OVERWRITE_PATCH)
class LULC(MultiValueEnum):
""" Enum class containing basic LULC types
"""
NO_DATA = 'No Data', 0, '#ffffff'
CULTIVATED_LAND = 'Cultivated Land', 1, '#ffff00'
FOREST = 'Forest', 2, '#054907'
GRASSLAND = 'Grassland', 3, '#ffa500'
SHRUBLAND = 'Shrubland', 4, '#806000'
WATER = 'Water', 5, '#069af3'
WETLAND = 'Wetlands', 6, '#95d0fc'
TUNDRA = 'Tundra', 7, '#967bb6'
ARTIFICIAL_SURFACE = 'Artificial Surface', 8, '#dc143c'
BARELAND = 'Bareland', 9, '#a6a6a6'
SNOW_AND_ICE = 'Snow and Ice', 10, '#000000'
@property
def id(self):
""" Returns an ID of an enum type
:return: An ID
:rtype: int
"""
return self.values[1]
@property
def color(self):
""" Returns class color
:return: A color in hexadecimal representation
:rtype: str
"""
return self.values[2]
def get_bounds_from_ids(ids):
bounds = []
for i in range(len(ids)):
if i < len(ids) - 1:
if i == 0:
diff = (ids[i + 1] - ids[i]) / 2
bounds.append(ids[i] - diff)
diff = (ids[i + 1] - ids[i]) / 2
bounds.append(ids[i] + diff)
else:
diff = (ids[i] - ids[i - 1]) / 2
bounds.append(ids[i] + diff)
return bounds
# Reference colormap things
lulc_bounds = get_bounds_from_ids([x.id for x in LULC])
lulc_cmap = ListedColormap([x.color for x in LULC], name="lulc_cmap")
lulc_norm = BoundaryNorm(lulc_bounds, lulc_cmap.N)
# takes some time due to the large size of the reference data
land_use_ref_path = os.path.join(DATA_FOLDER, 'land_use_10class_reference_slovenia_partial.gpkg')
land_use_ref = gpd.read_file(land_use_ref_path)
rasterization_task = VectorToRaster(land_use_ref, (FeatureType.MASK_TIMELESS, 'LULC'),
values_column='lulcid', raster_shape=(FeatureType.MASK, 'IS_DATA'),
raster_dtype=np.uint8)
# Define the workflow
workflow = LinearWorkflow(
add_data,
ndvi,
ndwi,
ndbi,
add_sh_valmask,
count_val_sh,
rasterization_task,
save
)
# Let's visualize it
workflow.dependency_graph()
SentinelHubInputTask NormalizedDifferenceIndexTask NormalizedDifferenceIndexTask_1 NormalizedDifferenceIndexTask_2 AddValidDataMaskTask CountValid VectorToRaster SaveTask
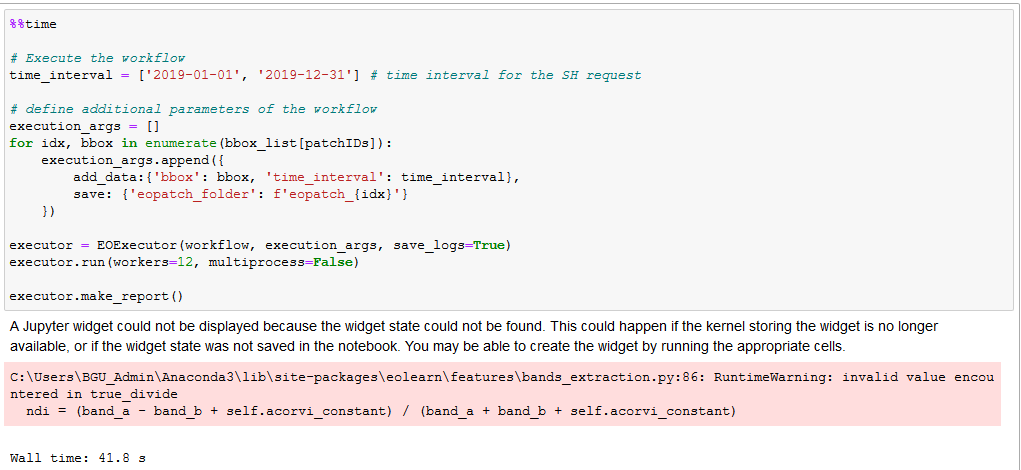
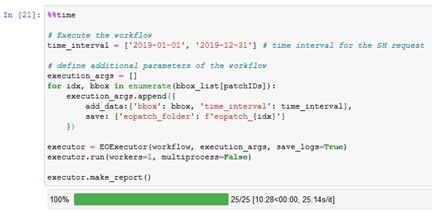
%%time
# Execute the workflow
time_interval = ['2019-01-01', '2019-12-31'] # time interval for the SH request
# define additional parameters of the workflow
execution_args = []
for idx, bbox in enumerate(bbox_list[patchIDs]):
execution_args.append({
add_data:{'bbox': bbox, 'time_interval': time_interval},
save: {'eopatch_folder': f'eopatch_{idx}'}
})
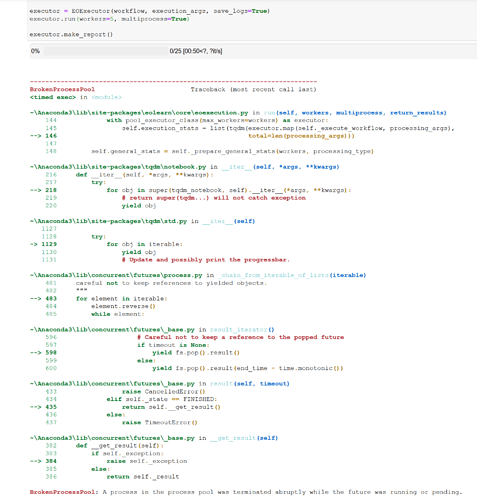
executor = EOExecutor(workflow, execution_args, save_logs=True)
executor.run(workers=12, multiprocess=False)
executor.make_report()
C:\Users\BGU_Admin\Anaconda3\lib\site-packages\eolearn\features\bands_extraction.py:86: RuntimeWarning: invalid value encountered in true_divide
ndi = (band_a - band_b + self.acorvi_constant) / (band_a + band_b + self.acorvi_constant)
Wall time: 39.6 s
__________________________________________________________________________________
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
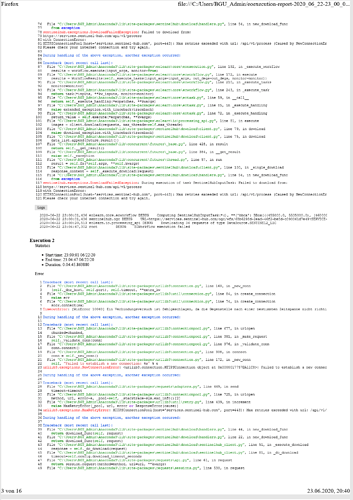
________________________________Error-file_____________________________________________
**Error-file:**
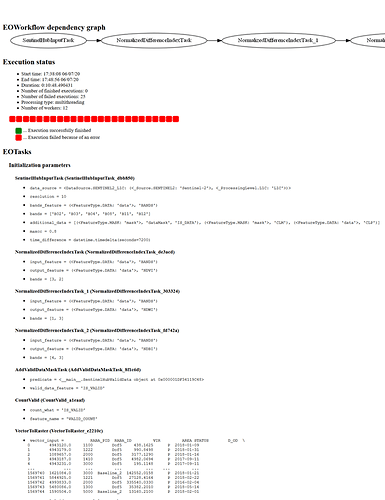
Execution status
Start time: 19:33:33 06/14/20
End time: 19:34:08 06/14/20
Duration: 0:00:35.149022
Number of finished executions: 0
Number of failed executions: 1
Processing type: multithreading
Number of workers: 12
... Execution successfully finished
... Execution failed because of an error
EOTasks
Initialization parameters
SentinelHubInputTask (SentinelHubInputTask_e35078)
data_source = <DataSource.SENTINEL2_L1C: (<_Source.SENTINEL2: 'Sentinel-2'>, <_ProcessingLevel.L1C: 'L1C'>)>
resolution = 10
bands_feature = (<FeatureType.DATA: 'data'>, 'BANDS')
bands = ['B02', 'B03', 'B04', 'B08', 'B11', 'B12']
additional_data = [(<FeatureType.MASK: 'mask'>, 'dataMask', 'IS_DATA'), (<FeatureType.MASK: 'mask'>, 'CLM'), (<FeatureType.DATA: 'data'>, 'CLP')]
maxcc = 0.8
time_difference = datetime.timedelta(seconds=7200)
NormalizedDifferenceIndexTask (NormalizedDifferenceIndexTask_1e4fb1)
input_feature = (<FeatureType.DATA: 'data'>, 'BANDS')
output_feature = (<FeatureType.DATA: 'data'>, 'NDVI')
bands = [3, 2]
NormalizedDifferenceIndexTask_1 (NormalizedDifferenceIndexTask_c0c30a)
input_feature = (<FeatureType.DATA: 'data'>, 'BANDS')
output_feature = (<FeatureType.DATA: 'data'>, 'NDWI')
bands = [1, 3]
NormalizedDifferenceIndexTask_2 (NormalizedDifferenceIndexTask_f962b4)
input_feature = (<FeatureType.DATA: 'data'>, 'BANDS')
output_feature = (<FeatureType.DATA: 'data'>, 'NDBI')
bands = [4, 3]
AddValidDataMaskTask (AddValidDataMaskTask_79928d)
predicate = <__main__.SentinelHubValidData object at 0x000001EA03892AC8>
valid_data_feature = 'IS_VALID'
CountValid (CountValid_4ed3eb)
count_what = 'IS_VALID'
feature_name = 'VALID_COUNT'
VectorToRaster (VectorToRaster_55e961)
vector_input = RABA_PID RABA_ID VIR AREA STATUS D_OD \
0 4943120.0 1100 Dof5 438.1625 P 2018-01-09
1 4943179.0 1222 Dof5 990.8498 P 2018-01-31
2 1089657.0 2000 Dof5 3177.1290 P 2018-01-16
3 4943187.0 1410 Dof5 4982.0694 P 2017-09-11
4 4943231.0 3000 Dof5 195.1148 P 2017-09-11
... ... ... ... ... ... ...
1569740 1621084.0 3000 Baseline_2 142552.0158 P 2018-01-21
1569741 5844925.0 1221 Dof5 27128.4164 P 2018-02-22
1569742 4993833.0 2000 Dof5 335540.0330 P 2016-02-04
1569743 5480086.0 1300 Dof5 35382.2010 P 2018-05-14
1569744 1590504.0 5000 Baseline_2 13160.2100 P 2018-02-01
lulcid lulcname \
0 1 cultivated land
1 1 cultivated land
2 2 forest
3 4 schrubland
4 8 artificial surface
... ... ...
1569740 8 artificial surface
1569741 1 cultivated land
1569742 2 forest
1569743 3 grassland
1569744 4 schrubland
geometry
0 MULTIPOLYGON (((394793.882 5040217.190, 394792...
1 MULTIPOLYGON (((394572.984 5040401.611, 394568...
2 MULTIPOLYGON (((417562.359 5124368.001, 417559...
3 MULTIPOLYGON (((394595.552 5040470.227, 394598...
4 MULTIPOLYGON (((394591.353 5040468.660, 394595...
... ...
1569740 MULTIPOLYGON (((437398.343 5131241.527, 437398...
1569741 MULTIPOLYGON (((446875.197 5073635.445, 446846...
1569742 MULTIPOLYGON (((540867.718 5095500.025, 540865...
1569743 MULTIPOLYGON (((528341.088 5138758.297, 528325...
1569744 MULTIPOLYGON (((400184.696 5147192.187, 400159...
[1569745 rows x 9 columns]
raster_feature = (<FeatureType.MASK_TIMELESS: 'mask_timeless'>, 'LULC')
SaveTask (SaveTask_15bd1e)
path = './eopatches/'
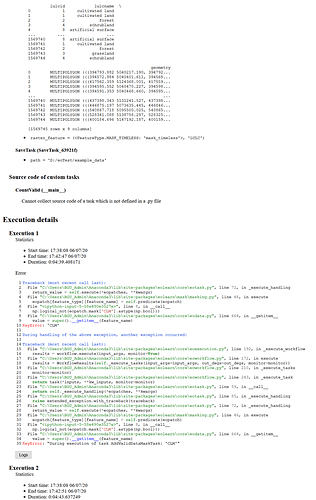
Source code of custom tasks
CountValid (__main__)
Cannot collect source code of a task which is not defined in a .py file
Execution details
Execution 1
Statistics
Start time: 19:33:33 06/14/20
End time: 19:34:08 06/14/20
Duration: 0:00:35.140987
Error
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
Traceback (most recent call last):
File "C:\Users\BGU_Admin\Anaconda3\lib\site-packages\eolearn\core\eotask.py", line 72, in _execute_handling
return_value = self.execute(*eopatches, **kwargs)
File "C:\Users\BGU_Admin\Anaconda3\lib\site-packages\eolearn\mask\masking.py", line 46, in execute
eopatch[feature_type][feature_name] = self.predicate(eopatch)
File "<ipython-input-36-58e490e3527e>", line 8, in __call__
np.logical_not(eopatch.mask['CLM'].astype(np.bool)))
File "C:\Users\BGU_Admin\Anaconda3\lib\site-packages\eolearn\core\eodata.py", line 664, in __getitem__
value = super().__getitem__(feature_name)
KeyError: 'CLM'
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "C:\Users\BGU_Admin\Anaconda3\lib\site-packages\eolearn\core\eoexecution.py", line 192, in _execute_workflow
results = workflow.execute(input_args, monitor=True)
File "C:\Users\BGU_Admin\Anaconda3\lib\site-packages\eolearn\core\eoworkflow.py", line 172, in execute
results = WorkflowResults(self._execute_tasks(input_args=input_args, out_degs=out_degs, monitor=monitor))
File "C:\Users\BGU_Admin\Anaconda3\lib\site-packages\eolearn\core\eoworkflow.py", line 210, in _execute_tasks
monitor=monitor)
File "C:\Users\BGU_Admin\Anaconda3\lib\site-packages\eolearn\core\eoworkflow.py", line 243, in _execute_task
return task(*inputs, **kw_inputs, monitor=monitor)
File "C:\Users\BGU_Admin\Anaconda3\lib\site-packages\eolearn\core\eotask.py", line 59, in __call__
return self._execute_handling(*eopatches, **kwargs)
File "C:\Users\BGU_Admin\Anaconda3\lib\site-packages\eolearn\core\eotask.py", line 85, in _execute_handling
raise extended_exception.with_traceback(traceback)
File "C:\Users\BGU_Admin\Anaconda3\lib\site-packages\eolearn\core\eotask.py", line 72, in _execute_handling
return_value = self.execute(*eopatches, **kwargs)
File "C:\Users\BGU_Admin\Anaconda3\lib\site-packages\eolearn\mask\masking.py", line 46, in execute
eopatch[feature_type][feature_name] = self.predicate(eopatch)
File "<ipython-input-36-58e490e3527e>", line 8, in __call__
np.logical_not(eopatch.mask['CLM'].astype(np.bool)))
File "C:\Users\BGU_Admin\Anaconda3\lib\site-packages\eolearn\core\eodata.py", line 664, in __getitem__
value = super().__getitem__(feature_name)
KeyError: "During execution of task AddValidDataMaskTask: 'CLM'"