Hello,
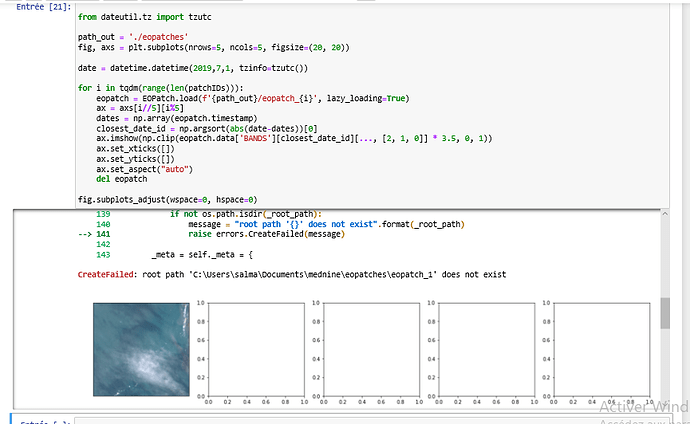
I have multiple areas (bboxes) and I want to prepare EOPatches, and each EOPatch should contain S2-L2A images for a set of dates (not date range). The problem is I can’t find a way using the eo-learn package to get S2 images for different dates at once, for example via SentinelHubInputTask.
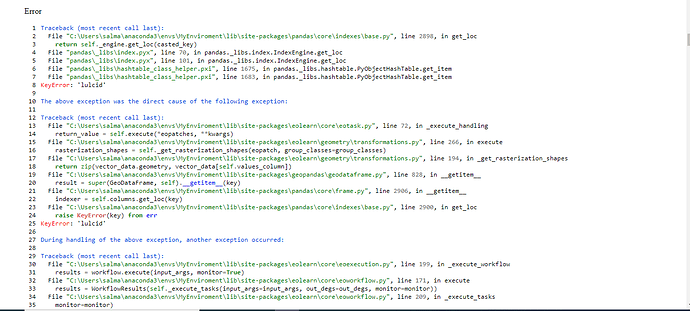
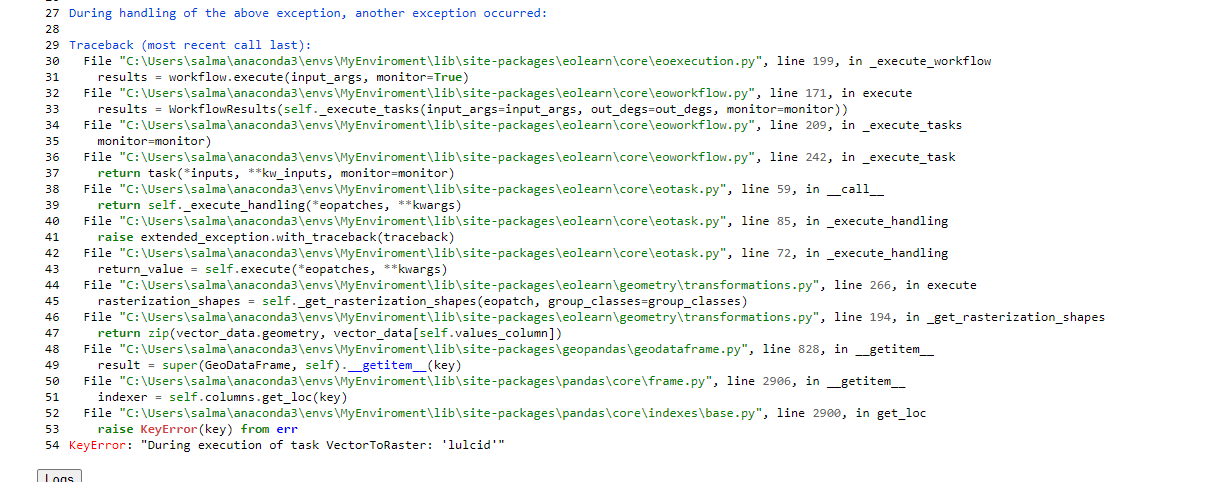
Using the code posted below, I get a ValueError: Trying to write data to an existing eopatch with a different timestamp.
Is there a way inside eo-learn to pack all S2 images (all bands) for selected dates into one EOPatch (i.a. FeatureType.DATA, ‘BANDS’) and then save it?
I also tried using EOExecutor with dates as different axecution arguments but, I was only able to get an EOPatch containing images for one date. It looks like the SaveTask overwrites EOPatch in every iteration of for loop.
Code that returns ValueError:
band_names = [‘B01’, ‘B02’, ‘B03’, ‘B04’, ‘B05’, ‘B06’, ‘B07’, ‘B08’, ‘B8A’, ‘B09’, ‘B11’, ‘B12’]
add_data = SentinelHubInputTask(
bands_feature=(FeatureType.DATA, ‘BANDS’),
bands = band_names,
resolution=10,
maxcc=0.8,
time_difference=datetime.timedelta(minutes=120),
data_source=DataSource.SENTINEL2_L2A
)
path_out = ‘./eopatches/’
save = SaveTask(path_out, overwrite_permission=OverwritePermission.OVERWRITE_FEATURES)
bbox=BBox(232087.50,2070306.50,232327.50,2070546.50), crs=CRS.UTM_16N)
dates=[‘2020-02-29’, ‘2020-4-04’, ‘2020-04-14’]
eopatch=EOPatch()
for date in dates:
add_data.execute(eopatch=eopatch, bbox=bbox, time_interval= date)
save.execute(eopatch=eopatch, eopatch_folder=‘eopatch_selected_dates’)