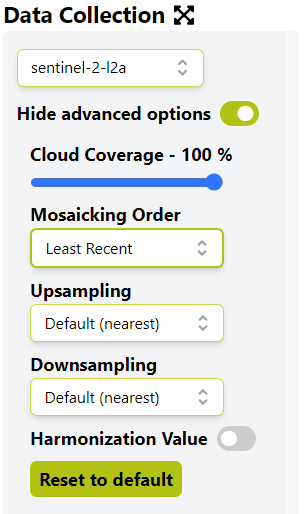
@megha.devaraju please find the curl-command posted below. in the example below, the satellite time range is set as follows:
"data": [
{
"dataFilter": {
"timeRange": {
"from": "2023-03-12T00:00:00Z",
"to": "2023-08-07T23:59:59Z"
}
},
"type": "sentinel-2-l2a"
}
]
when i change the time range to, for example, "from": "2022-03-12T00:00:00Z", "to": "2023-08-07T23:59:59Z"
i receive two identical tiffd
would you please help me solve this issue?
curl-command
curl -X POST https://services.sentinel-hub.com/api/v1/process \
-H 'Content-Type: application/json' \
-H 'Authorization: Bearer xxx' \
-H 'Accept: application/tar' \
-d '{
"input": {
"bounds": {
"geometry": {
"type": "Polygon",
"coordinates": [
[
[
1579250.82367497,
6861560.75494123
],
[
1579262.05009723,
6861568.0993758
],
[
1579274.53666044,
6861572.98902966
],
[
1579287.75641312,
6861575.2175475
],
[
1579301.15146001,
6861574.6908805
],
[
1579314.15650793,
6861571.431256
],
[
1580614.32240361,
6861098.63910992
],
[
1580626.20934827,
6861092.89222752
],
[
1580636.69971985,
6861084.86506323
],
[
1580645.36441237,
6861074.88597658
],
[
1580651.8490035,
6861063.36316948
],
[
1580655.88825164,
6861050.76798707
],
[
1580657.31694429,
6861037.61563564
],
[
1580656.07665425,
6861024.44410741
],
[
1580652.2181275,
6861011.79217384
],
[
1580213.19772315,
6859964.91889461
],
[
1580207.16807032,
6859953.72991754
],
[
1580199.09260609,
6859943.92221921
],
[
1580189.27663079,
6859935.86657521
],
[
1580178.09124157,
6859929.86752367
],
[
1580165.95930318,
6859926.15185373
],
[
1580153.33946234,
6859924.86003288
],
[
1580140.70880969,
6859926.0408975
],
[
1580128.5448448,
6859929.64980675
],
[
1578507.57893836,
6860588.16085359
],
[
1578495.82259782,
6860594.40233978
],
[
1578485.57332546,
6860602.90455025
],
[
1578477.25717289,
6860613.31406842
],
[
1578471.21983892,
6860625.19819229
],
[
1578467.71229861,
6860638.06291925
],
[
1578466.88036911,
6860651.37347985
],
[
1578468.75864619,
6860664.57656646
],
[
1578473.26906401,
6860677.12333377
],
[
1578480.22413819,
6860688.49221449
],
[
1578750.38594929,
6861043.08417387
],
[
1578750.45489957,
6861043.17450606
],
[
1578969.96138274,
6861330.22514622
],
[
1578980.66035828,
6861341.24507847
],
[
1579250.82367497,
6861560.75494123
]
]
]
},
"properties": {
"crs": "http://www.opengis.net/def/crs/EPSG/0/3857"
}
},
"data": [
{
"dataFilter": {
"timeRange": {
"from": "2023-03-12T00:00:00Z",
"to": "2023-08-07T23:59:59Z"
}
},
"type": "sentinel-2-l2a"
}
]
},
"output": {
"width": 512,
"height": 276.639,
"responses": [
{
"identifier": "default",
"format": {
"type": "image/tiff"
}
},
{
"identifier": "userdata",
"format": {
"type": "application/json"
}
}
]
},
"evalscript": "//VERSION=3\n// Script to extract a time series of NDVI values using \n// Sentinel 2 Level 2A data and metadata file.\nfunction setup() {\n return {\n input: [{\n bands: [\"B04\", \"B08\"],\n units: \"DN\"\n }],\n output: {\n bands: 1,\n sampleType: SampleType.FLOAT32\n },\n mosaicking: Mosaicking.ORBIT\n }\n \n }\n \n // The following function is designed to update the number of\n // output bands without knowing beforehand how many there are\n function updateOutput(outputs, collection) {\n Object.values(outputs).forEach((output) => {\n output.bands = collection.scenes.length;\n });\n }\n // function to generate a json file with a list of the NDVI \n // dates used in the analysis. \n function updateOutputMetadata(scenes, inputMetadata, outputMetadata) {\n var dds = [];\n for (i=0; i<scenes.length; i++){\n dds.push(scenes[i].date)\n }\n outputMetadata.userData = { \"acquisition_dates\": JSON.stringify(dds) }\n }\n \n function evaluatePixel(samples) {\n // Precompute an array to contain NDVI observations\n var n_observations = samples.length;\n let ndvi = new Array(n_observations).fill(0);\n \n // Fill the array with NDVI values\n samples.forEach((sample, index) => {\n ndvi[index] = (sample.B08 - sample.B04) / (sample.B08 + sample.B04) ;\n });\n \n return ndvi;\n }"
}'